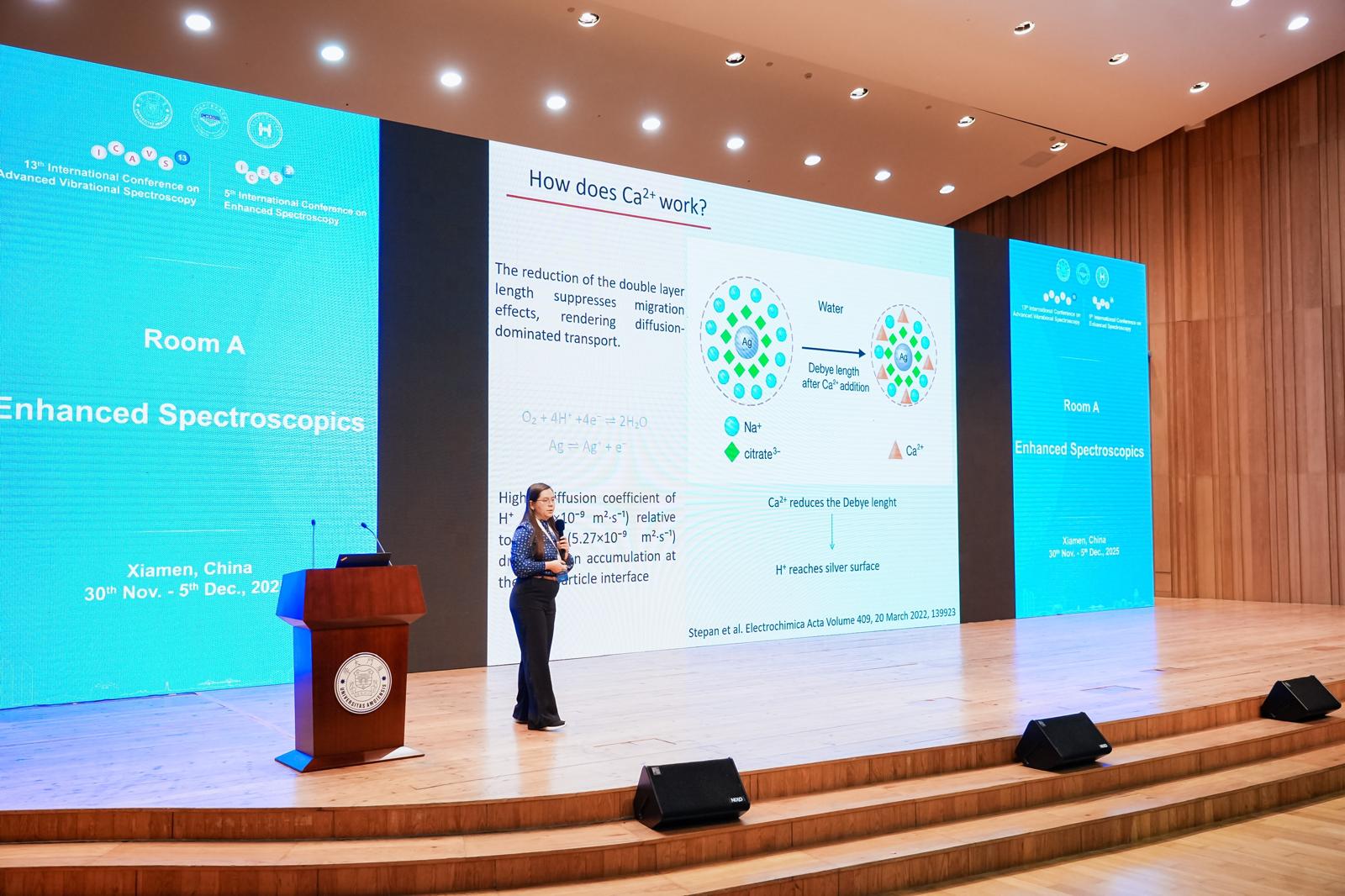
The first stage of the project involved obtaining DNA fragments by PCR amplification, generating sequences with different degrees of methylation (192 bp in length using the SEPT9 gene-specific primer with methylation levels of 0.54% and 8.89%, respectively). Based on these fragments, SERS detection was subsequently optimized by identifying the appropriate metallic substrate. Silver colloids obtained by reduction with sodium borohydride, citrate, and hydroxylamine hydrochloride were compared, with the latter demonstrating the best SERS signal enhancement for the DNA fragments even at quantities as low as 5 ng.
Project Results
Scientific Results
For the optimal substrate (hya-AgNPs), various cations that facilitate DNA adsorption on the nanoparticle surface were tested (Ca2+, Mg2+, Zn2+, Cu2+, Al3+, Na+, and K+). Among these, Ca2+ generated the most intense SERS signal from the fragmented DNA, although fluorescence analysis did not indicate a significantly higher amount adsorbed compared to the other cations. This result suggests that the role of Ca2+ is not limited to facilitating adsorption but also contributes to an increased overall enhancement factor. Thus, Ca2+ was identified as the optimal ion for SERS analysis of DNA fragments, and hya-AgNPs as the preferred metallic substrate. A change in the SERS signal was also observed depending on the DNA concentration, with DNA amounts below 10 ng being ideal for SERS analysis.
Subsequently, the extracellular DNA extraction protocol from the culture medium of malignant and benign cell lines was optimized. It was demonstrated that collecting the medium after 7 days of incubation leads to a significant increase in the amount of recovered DNA. For 15 ml culture medium volumes, lyophilization of the sample followed by redispersion in 1 ml of ultrapure water before the extraction step was necessary to ensure sufficient concentration for further analyses. For malignant cell lines (MDAMB231, SKHEP1, NB4), the extracellular DNA concentration extracted was 34–63 ng/µl, while for non-tumor cell lines (CCD1137Sk, HS27) it was approximately 10–17 ng/µl.
Articles
- S. D. Iancu et al., “SERS-based detection of DNA methylation for cancer diagnosis: Cation-mediated adsorption to silver nanoparticles,” PloS one, vol. 20, no. 6, p. e0325539, 2025.
- A. M. Chiriac et al., “Citrate-reduced silver nanoparticles: Synthesis temperature dependent properties,” Applied Surface Science, p. 163759, 2025.
- D. Andras et al., "SERS liquid biopsy in colorectal cancer detection and treatment response: Revealing metabolic memory post-radiochemotherapy", Nanomedicine: Nanotechnology, Biology and Medicine, Volume 68, August 2025, 102840.
- D. Andras et al., "Advancing Breast Cancer Diagnosis: Optimization of Raman Spectroscopy for Urine-Based Early Detection", Biomedicines 2025, 13(2), 505.
Presentations
21th International Conference "Students for Students", Cluj-Napoca, 9th-13th April 2025
Author: G. Ion
Title: Label-Free SERS Detection of Glucose Using Glucose-Reduced Silver Nanoparticles
Women in Photonics, Jena, Germany, 1-5 June 2025
Author: S. D. Iancu
Title: Nanoparticle-Enhanced SERS for Rapid Detection of DNA Methylation for Cancer Diagnostics




